
Welcome to the help files of
Part B - The different type of results
Three different types of outputs are avalaible for the results:
- short results, with only the sequence and the prediction in terms of Protein Blocks.
- normal results, with the sequence, the prediction in terms of Protein Blocks and an index
derived from the entropy-based index Neq to evaluate the results of prediction.
- complete results, with amino acid sequence, the local structure prediction
in terms of Protein Blocks
and the whole Protein Blocks with their associated probabilities.
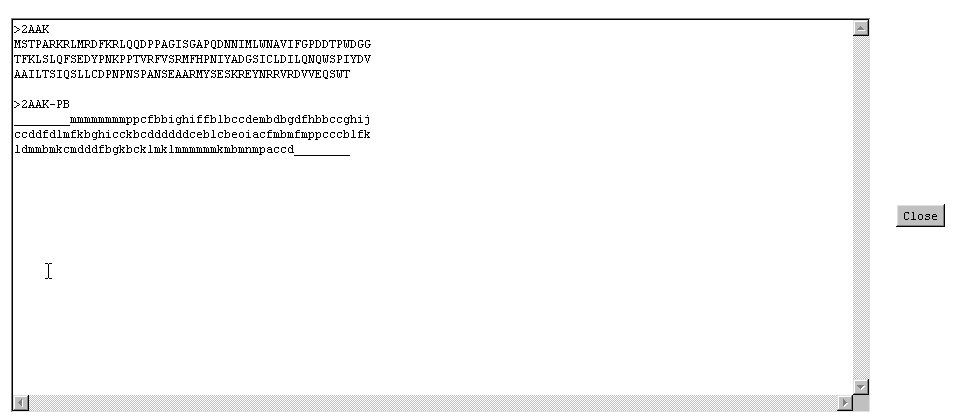
- sequence
- the sequence in amino acids.
- prediction
- the prediction in terms of Protein Blocks with only the most probable Protein Blocks.
Top
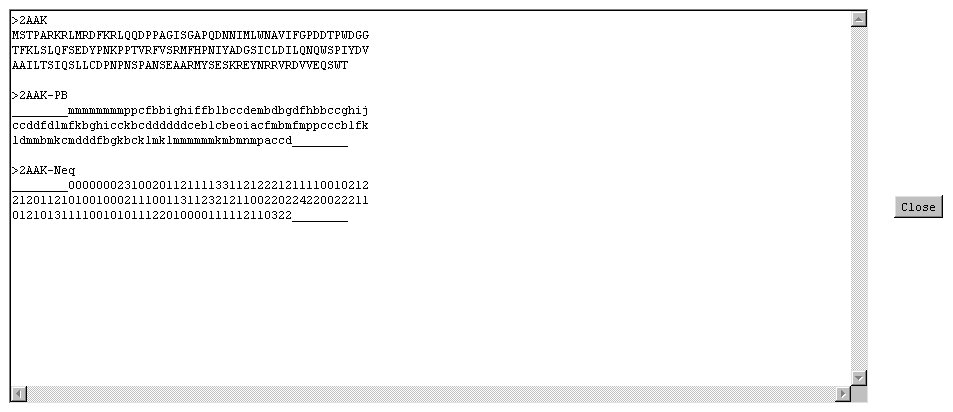
- sequence
- the sequence in amino acids.
- prediction
- the prediction in terms of Protein Blocks with only the most probable Protein Blocks.
- normalised Neq
- the entropy-based index Neq varies between 1 (sequence of high informativity)
and 16 (sequence without informativity). Warning: the scale has been
normalized - for clarity purpose - in the range [0:9] instead of the classical [1:16].
Top
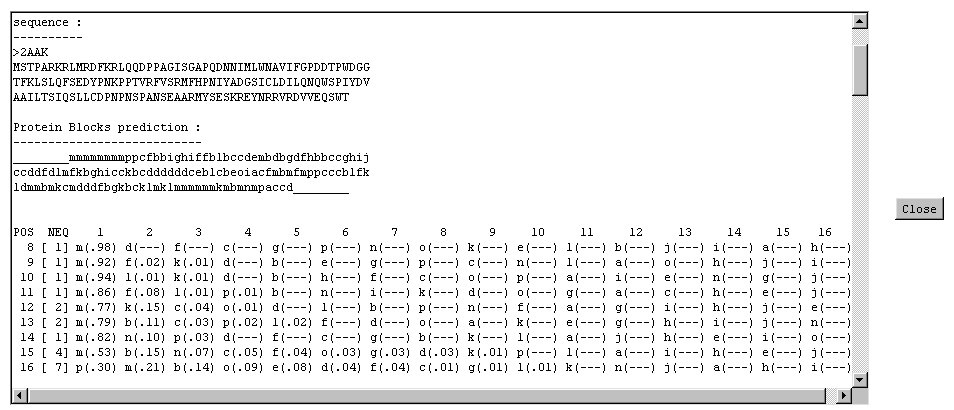

- sequence
- the sequence in amino acids.
- prediction
- the prediction in terms of Protein Blocks with only the most probable Protein Blocks.
- complete prediction
- the prediction with all the informations, with (from left to right) the position of the residue in the sequence,
the local Neq (in the range [1:16]) and then by decreasing order the 16 Protein Blocks with
their associated probability (the sum of all probability is 1, the symbol --- is for probability
less than 0.01).
back
Last modif : 25 April 2004